BirdDMD¶
Dynamic Mode Decomposition for bird flight analysis.
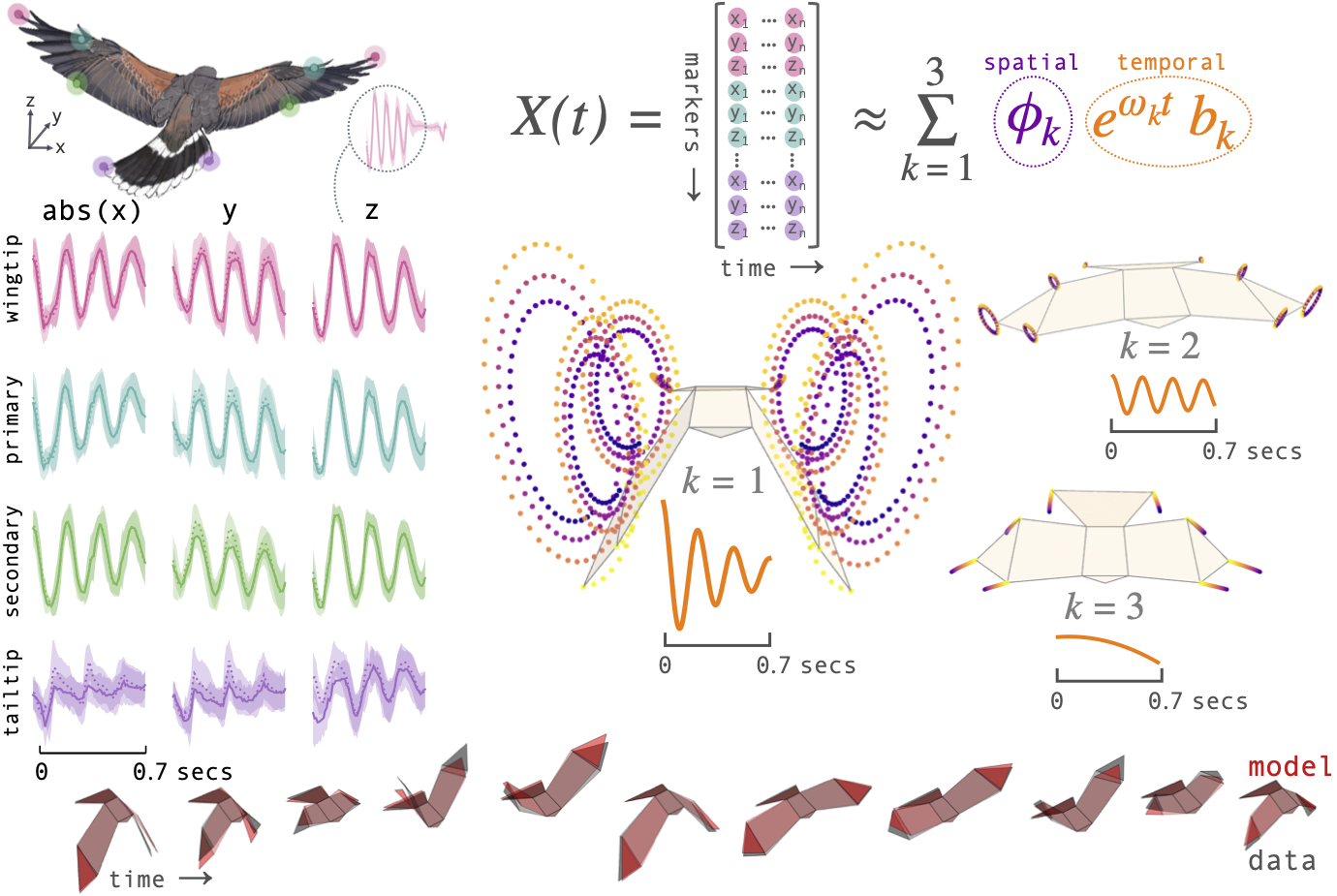
BirdDMD is a minimal, scientist-friendly toolkit for decomposing motion-capture time series into spatially coherent oscillatory modes. It wraps PyDMD with sensible defaults for biological motion data and provides hawk-specific convenience functions built on the morphing_birds library for animation.
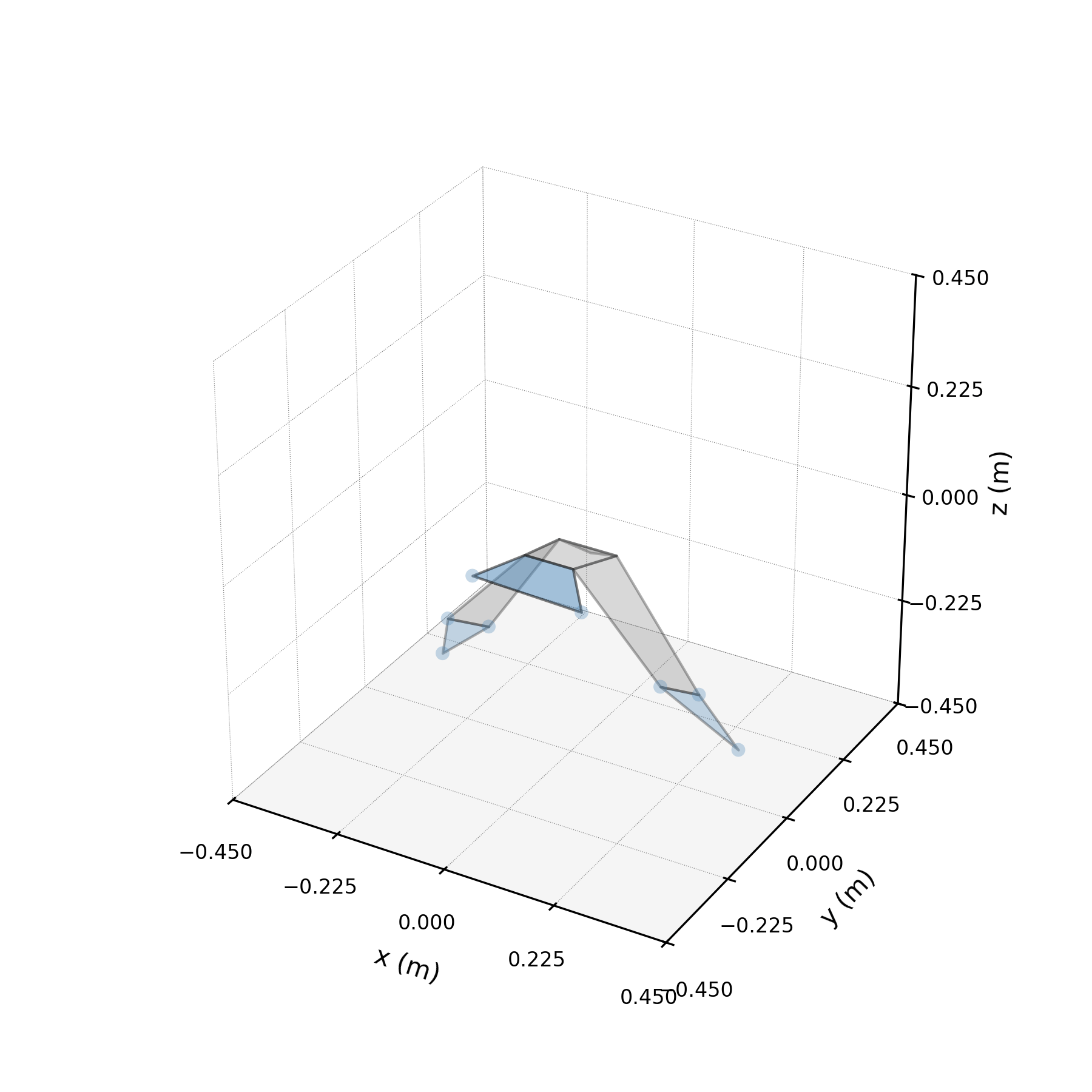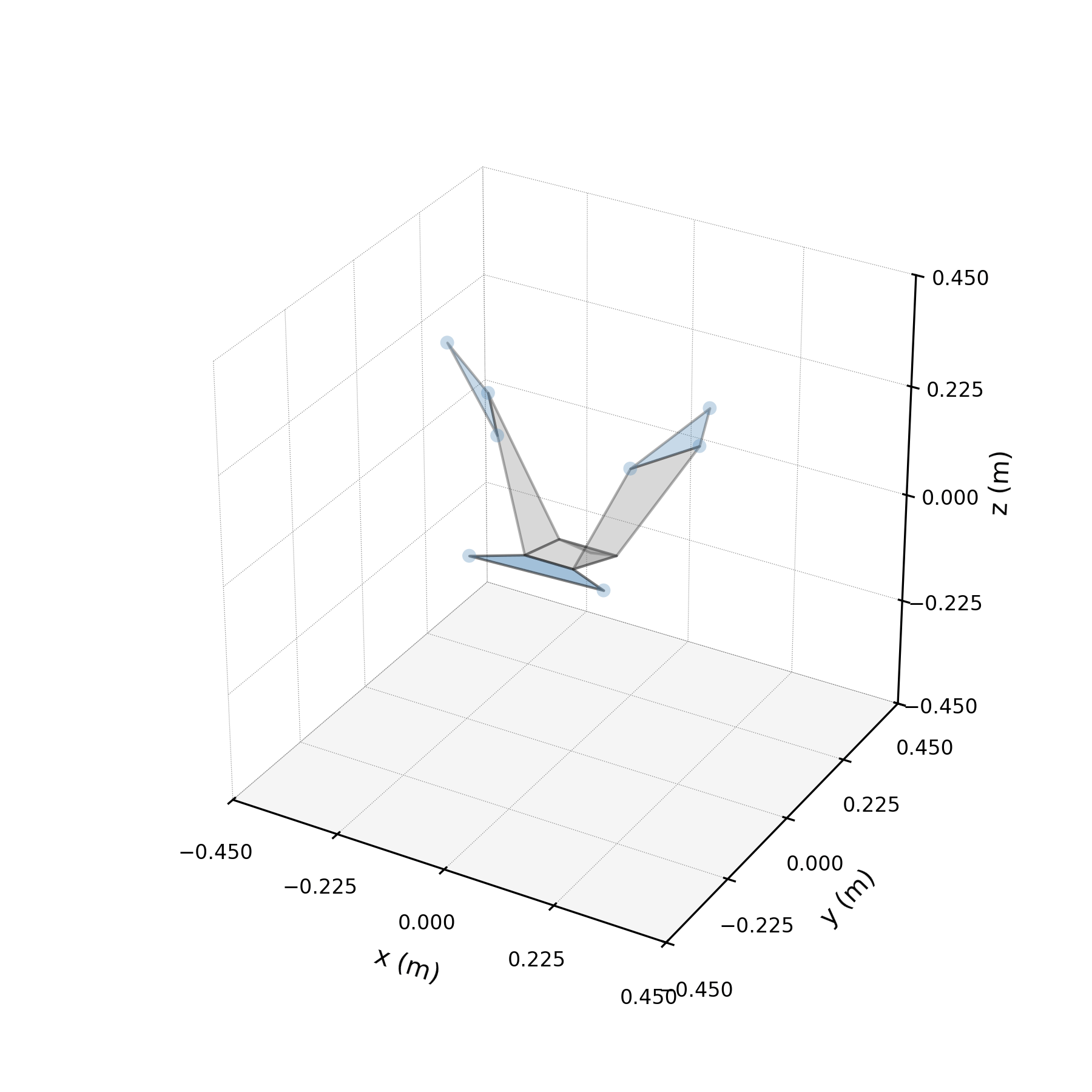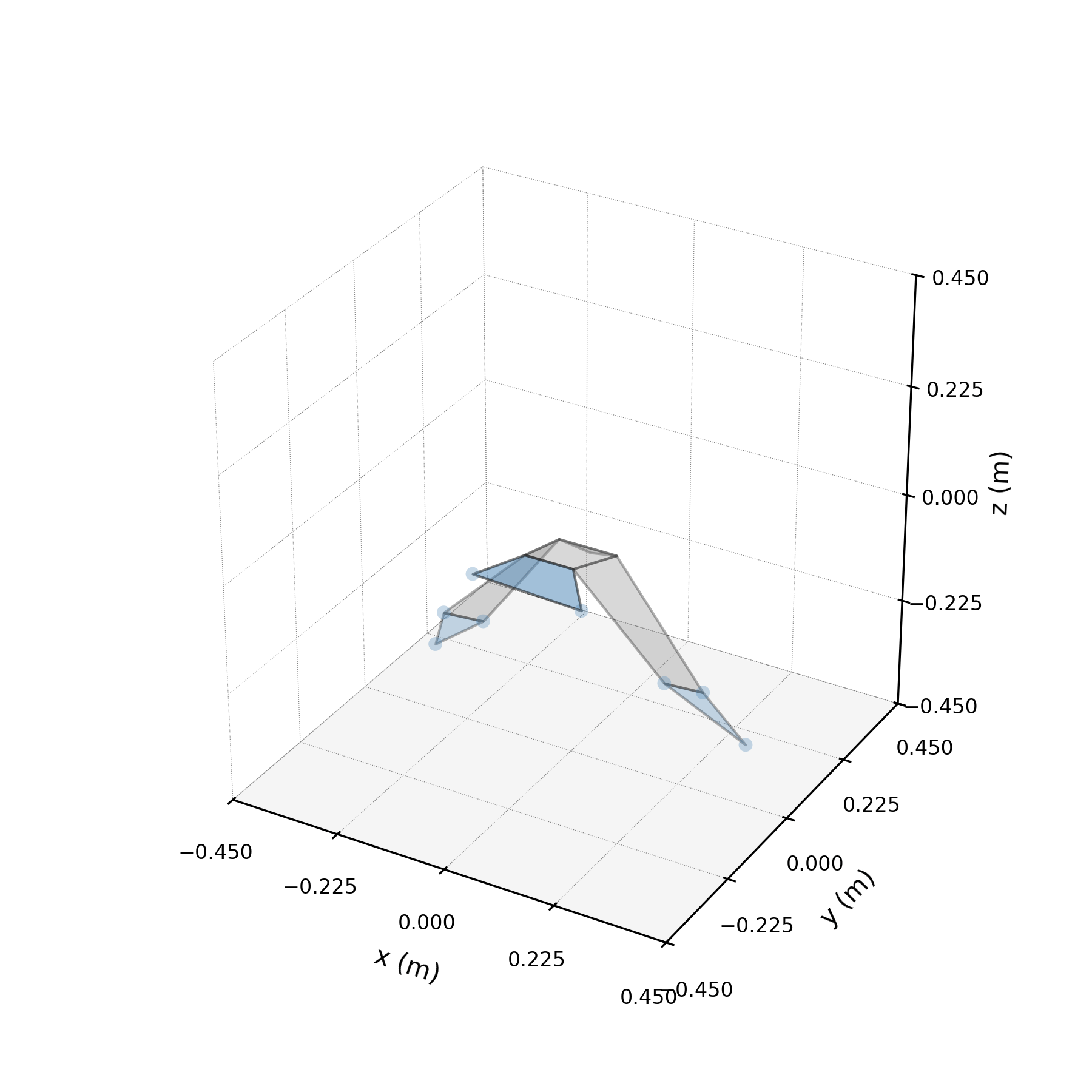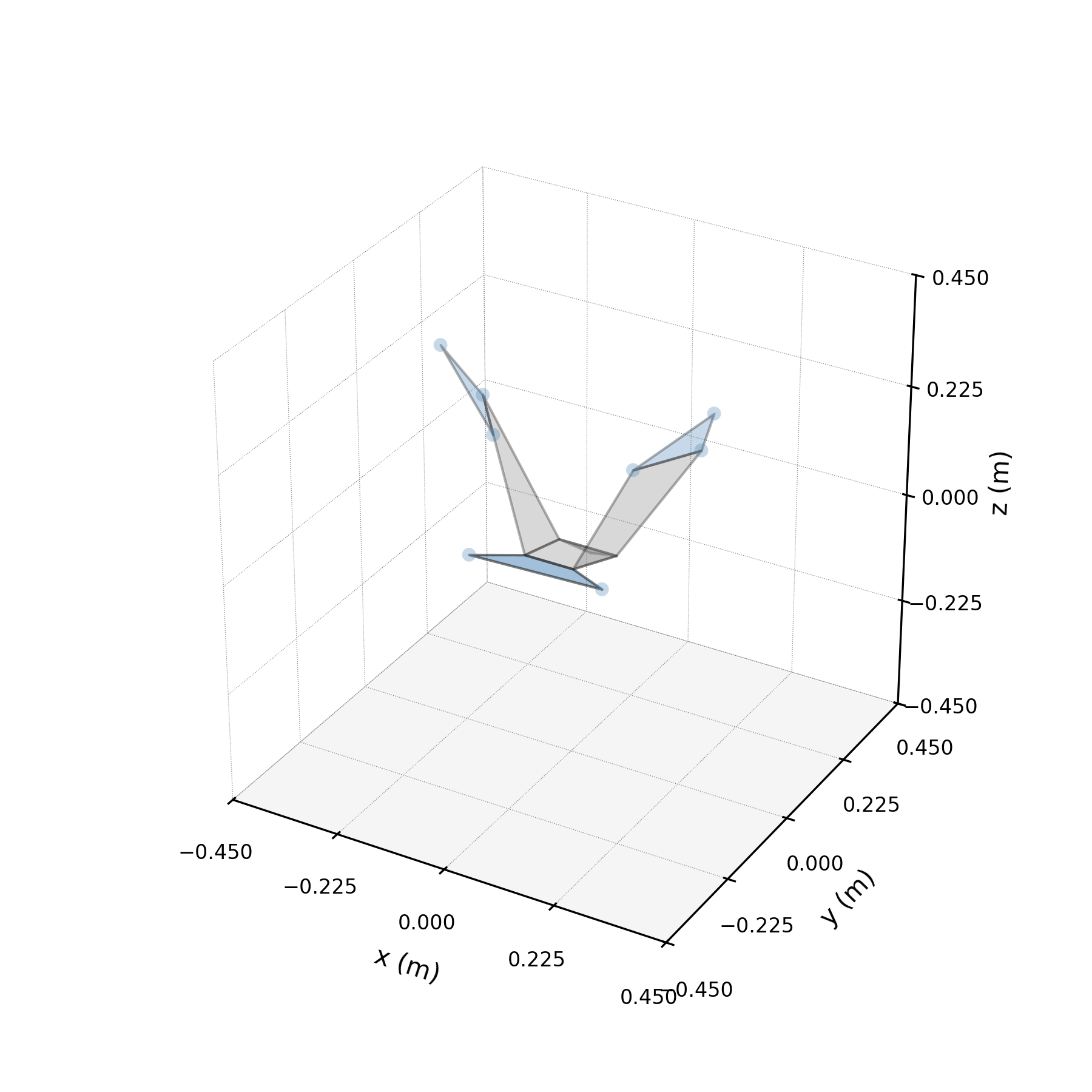
Architecture¶
BirdDMD
├── core ← run_dmd(), DMDResult, convergence analysis
├── reconstruction← reconstruct(), forecast()
├── data ← validation, normalisation, binning, reshaping
├── stats ← RMSE, variance explained, sequence filtering
├── plotting ← dataset-agnostic DMD visualisation
├── hawk ← hawk-specific loading, wrappers, and plots
└── _constants ← named constants (marker positions, colours)
The package separates dataset-agnostic DMD utilities (core, data,
reconstruction, stats, plotting) from hawk-specific functions
(hawk, _constants), so the core analysis can be reused with other
motion-capture datasets.
Quick links¶
- Quickstart guide — installation and first analysis
- API Reference — full module documentation
- Notebook Gallery — worked examples from the manuscript
Notebooks¶
| Notebook | Topic |
|---|---|
| 00 Introduction | DMD theory, synthetic example, and data description |
| 01 Flapping Modes | Flapping mode decomposition |
| 03 Double Frequency | The doubled-frequency mode (2ω) |
| 04 Full Flight | Full flight trajectory: flapping to gliding |
| 05 Turning | Obstacle avoidance and turning |
| 06 Reconstruction Accuracy | Batch RMSE statistics |
| 07 Generative Model | Forecasting, frequency modification, and synthetic flight |
Standard settings¶
- 8 bilateral markers × 3 axes = 24 coordinates per frame
n_modes=6,d=2,eig_constraints={"conjugate_pairs"}