07 — Generative model: forecasting and synthetic flight¶
DMD eigenvalues encode both oscillation frequency (imaginary part) and growth/decay rate (real part). By manipulating these eigenvalues we can use the fitted model generatively:
- Stabilised forecast — set the real part of each eigenvalue to zero, removing exponential growth or decay, to produce an indefinitely periodic prediction (Eq. 2 in the manuscript).
- Frequency modification — change the imaginary part to slow down or speed up specific modes while preserving their spatial structure.
- Synthetic flight generation — combine arbitrary subsets of modes over extended time ranges to create novel flight sequences.
- Temporal upsampling — evaluate the continuous-time model on a denser time grid to smoothly interpolate between observed frames.
%load_ext autoreload
%autoreload 2
%matplotlib widget
%config InlineBackend.figure_format='svg'
import matplotlib.pyplot as plt
import numpy as np
from morphing_birds import Hawk3D, animate_compare, animate_plotly_compare
from birddmd import (
expand_marker_sequence,
expand_time_sequence,
forecast,
reconstruct,
run_dmd,
)
1 — Fit DMD on flapping data¶
We re-fit DMD to the same Toothless 9 m binned flapping data used in 01_flapping_modes. The six modes (three conjugate pairs) provide the basis for all generative experiments below.
The preparation script (scripts/prepare_flapping.py) bins the raw wingbeat sequences. Expand the cell below to review the pipeline.
# %load scripts/prepare_flapping.py
loaded_data = np.load("../data/processed/Toothless_flapping_binned.npz")
markers, times = loaded_data["markers"], loaded_data["times"]
n_markers = 8
hawk3d = Hawk3D("../data/mean_hawk_shape.csv")
# ani = animate_compare(
# hawk3d, [markers],
# colour="blue", az=-60, el=30,
# )
# ani.save("figures/07_real_flight.gif", writer='Pillow', fps=40, dpi=300)
# print("Saved GIF: figures/07_real_flight.gif")
hawk3d = Hawk3D("../data/mean_hawk_shape.csv")
average_shape = hawk3d.markers
result = run_dmd(
data=markers,
times=times,
n_modes=6,
d=2,
eig_constraints={"conjugate_pairs"},
n_markers=n_markers,
average_shape=average_shape,
verbose=True,
)
Running DMD with 6 modes, delay d=2 Input shape: (139, 24) Number of variables in DMD results: 48 Complex Eigenvalues (log): [-0.813+28.32j -0.813-28.32j -0.228+55.056j -0.228-55.056j -0.887 -0.j -0.887 +0.j ] Number of modes in Psi: 6
2 — Growth/decay interpretation¶
Eigenvalues with negative real part decay over time; positive real part means exponential growth (physically unrealistic for long forecasts).
eigs = result._dmd_object.eigs
freqs = np.imag(eigs) / (2 * np.pi)
print("Eigenvalue analysis:")
print(f"{'Mode':>5} {'Real':>10} {'Freq (Hz)':>10} {'Growth/decay':>15}")
print("-" * 45)
for k in range(len(eigs)):
behaviour = (
"growth" if eigs[k].real > 0 else "decay" if eigs[k].real < 0 else "stable"
)
print(f"{k:5d} {eigs[k].real:10.4f} {freqs[k]:10.2f} {behaviour:>15}")
Eigenvalue analysis:
Mode Real Freq (Hz) Growth/decay
---------------------------------------------
0 -0.8129 4.51 decay
1 -0.8129 -4.51 decay
2 -0.2275 8.76 decay
3 -0.2275 -8.76 decay
4 -0.8866 -0.00 decay
5 -0.8866 0.00 decay
# ── Robust mode identification by frequency ─────────────────────────────
# Instead of relying on pair ordering (which has caused mix-ups before),
# identify each conjugate pair by its characteristic frequency.
conjugate_pairs = result.conjugate_pairs
pair_freq_hz = [result.pair_frequency(p) for p in range(result.n_pairs)]
WINGBEAT_PAIR = min(
range(len(conjugate_pairs)), key=lambda k: abs(pair_freq_hz[k] - 4.5)
)
FOLDING_PAIR = min(
range(len(conjugate_pairs)), key=lambda k: abs(pair_freq_hz[k] - 8.8)
)
BASE_PAIR = min(range(len(conjugate_pairs)), key=lambda k: abs(pair_freq_hz[k] - 0.0))
ALL_PAIRS = list(range(len(conjugate_pairs)))
# Convenience: raw mode indices for functions that need them
WINGBEAT_MODES = list(conjugate_pairs[WINGBEAT_PAIR])
FOLDING_MODES = list(conjugate_pairs[FOLDING_PAIR])
BASE_MODES = list(conjugate_pairs[BASE_PAIR])
print(
f"Wingbeat pair {WINGBEAT_PAIR}:"
f" modes {WINGBEAT_MODES} ({pair_freq_hz[WINGBEAT_PAIR]:.1f} Hz)"
)
print(
f"Folding pair {FOLDING_PAIR}:"
f"modes {FOLDING_MODES} ({pair_freq_hz[FOLDING_PAIR]:.1f} Hz)"
)
print(
f"Base pair {BASE_PAIR}:modes {BASE_MODES} ({pair_freq_hz[BASE_PAIR]:.1f} Hz)"
)
Wingbeat pair 0: modes [0, 1] (4.5 Hz) Folding pair 1:modes [2, 3] (8.8 Hz) Base pair 2:modes [4, 5] (0.0 Hz)
3 — Stabilised reconstruction (Eq. 2)¶
By setting include_growth_decay=False we use only the imaginary part of the eigenvalues, producing a stable periodic forecast that neither grows nor decays. This corresponds to Eq. 2 in the manuscript. Below we compare the stabilised forecast against the original (decaying) reconstruction, extending the time axis to 3× the training duration.
Only the two oscillatory conjugate pairs (wingbeat and harmonic) are used for forecasting — the base posture is already provided by average_shape, while pair 2 is a rapidly decaying transient that would freeze as a constant offset in the stabilised version.
fake_time = np.linspace(times[0], times[-1] * 3, len(times) * 3)
# With growth/decay (may diverge) — oscillatory pairs only
forecast_with_gd = forecast(
result,
fake_time,
pairs=[WINGBEAT_PAIR, FOLDING_PAIR],
stable=False,
)
# Stabilised (no growth/decay) — oscillatory pairs only
forecast_stable = forecast(
result,
fake_time,
pairs=[WINGBEAT_PAIR, FOLDING_PAIR],
stable=True,
)
# Compare wingtip z
fig, ax = plt.subplots(figsize=(10, 3))
ax.plot(fake_time, forecast_with_gd[:, 0, 2], label="With growth/decay")
ax.plot(fake_time, forecast_stable[:, 0, 2], label="Stabilised")
ax.axvline(times[-1], color="grey", ls=":", label="End of training data")
ax.set_xlabel("Time (s)")
ax.set_ylabel("Wingtip z")
ax.legend()
ax.set_title("Stabilised vs unstabilised forecast")
plt.tight_layout()
fig.savefig("figures/07_stabilised_forecast.svg", format="svg")
plt.show()
# hawk3d_forecast = Hawk3D("../data/mean_hawk_shape.csv")
# min_len = min(forecast_stable.shape[0], forecast_with_gd.shape[0])
# ani = animate_compare(
# hawk3d_forecast, [forecast_stable[:min_len], forecast_with_gd[:min_len]],
# colour="blue", az=-60, el=30,
# )
# ani.save("figures/07_stable_vs_unstable.gif", writer='Pillow', fps=40, dpi=300)
# print("Saved GIF: figures/07_stable_vs_unstable.gif")
3b — Extended forecast¶
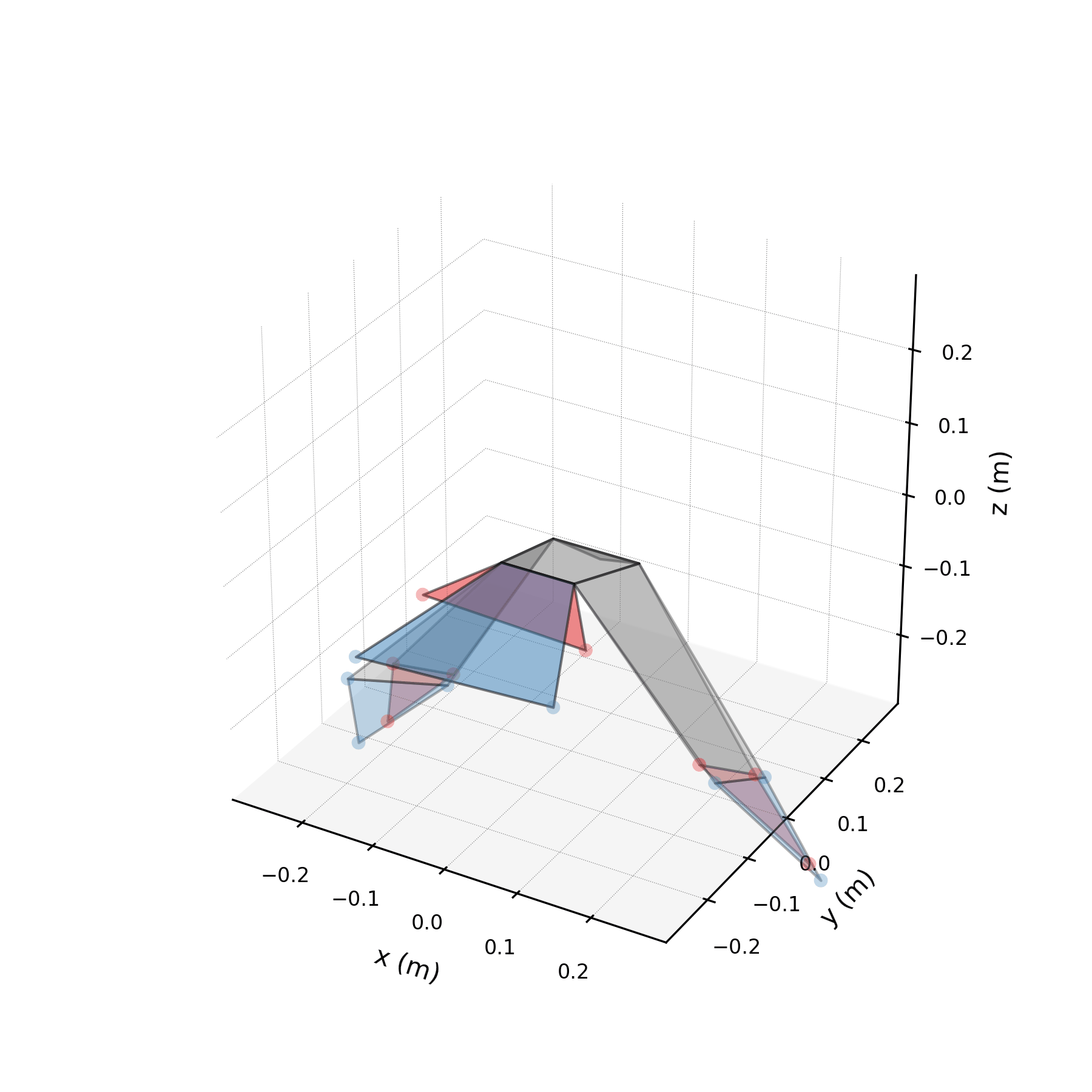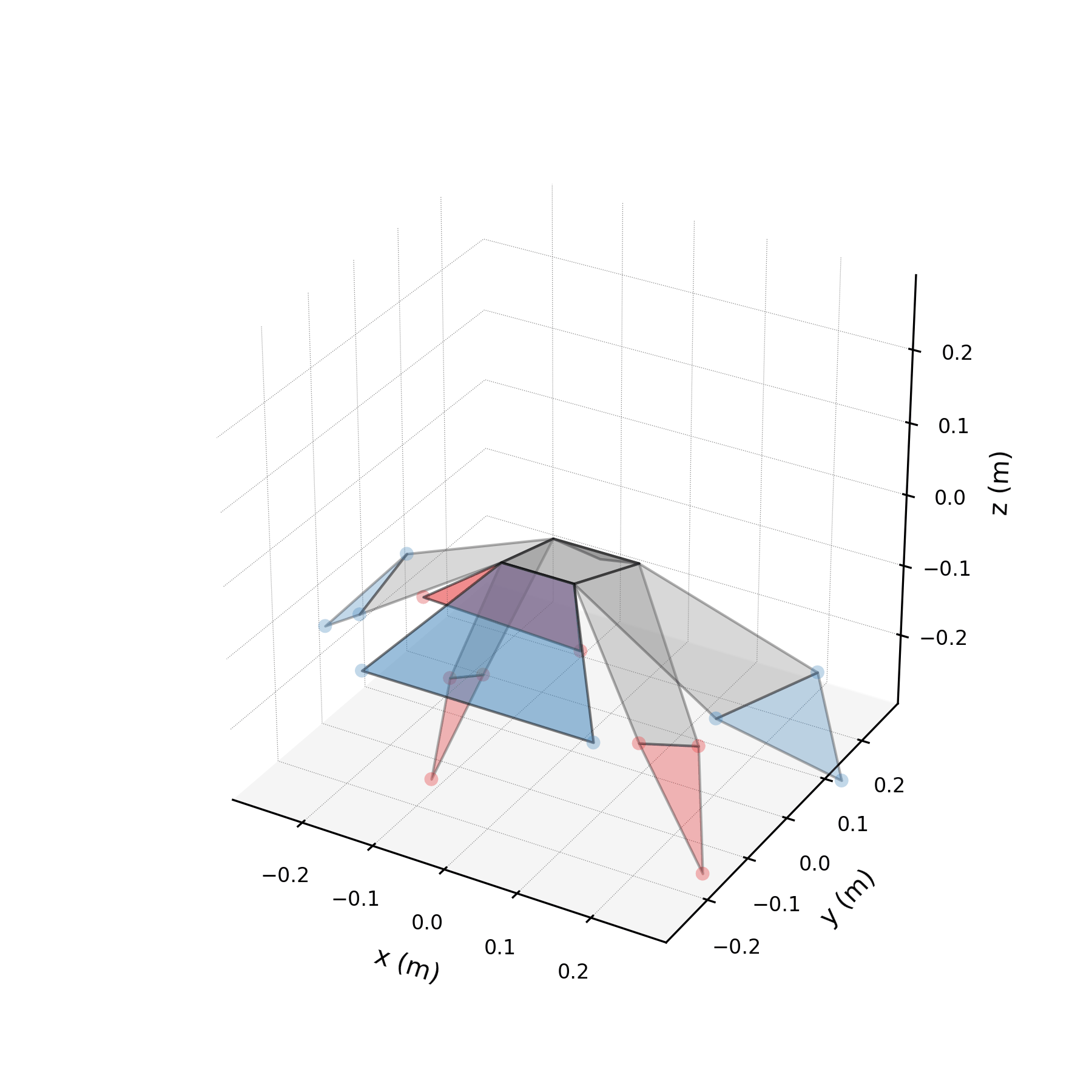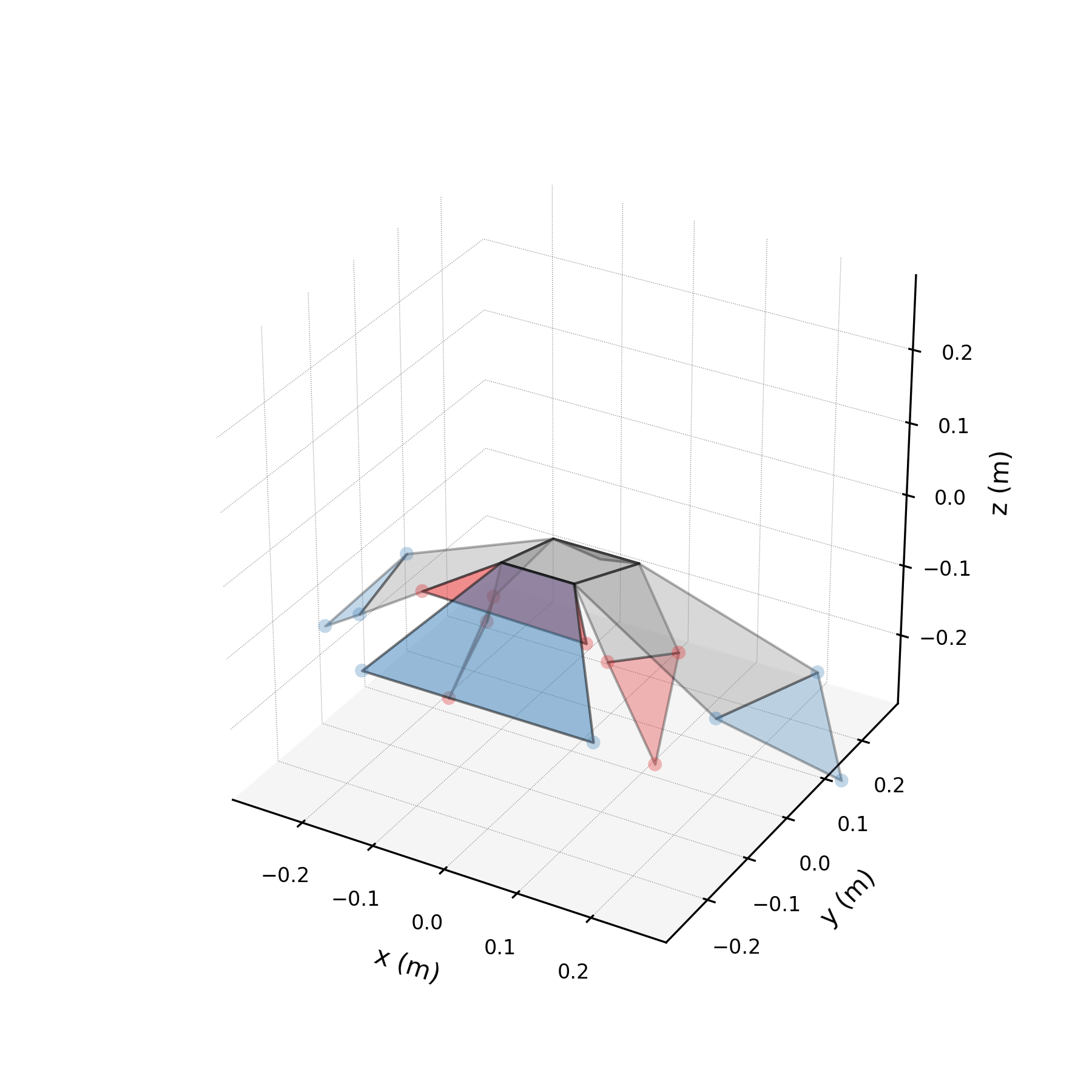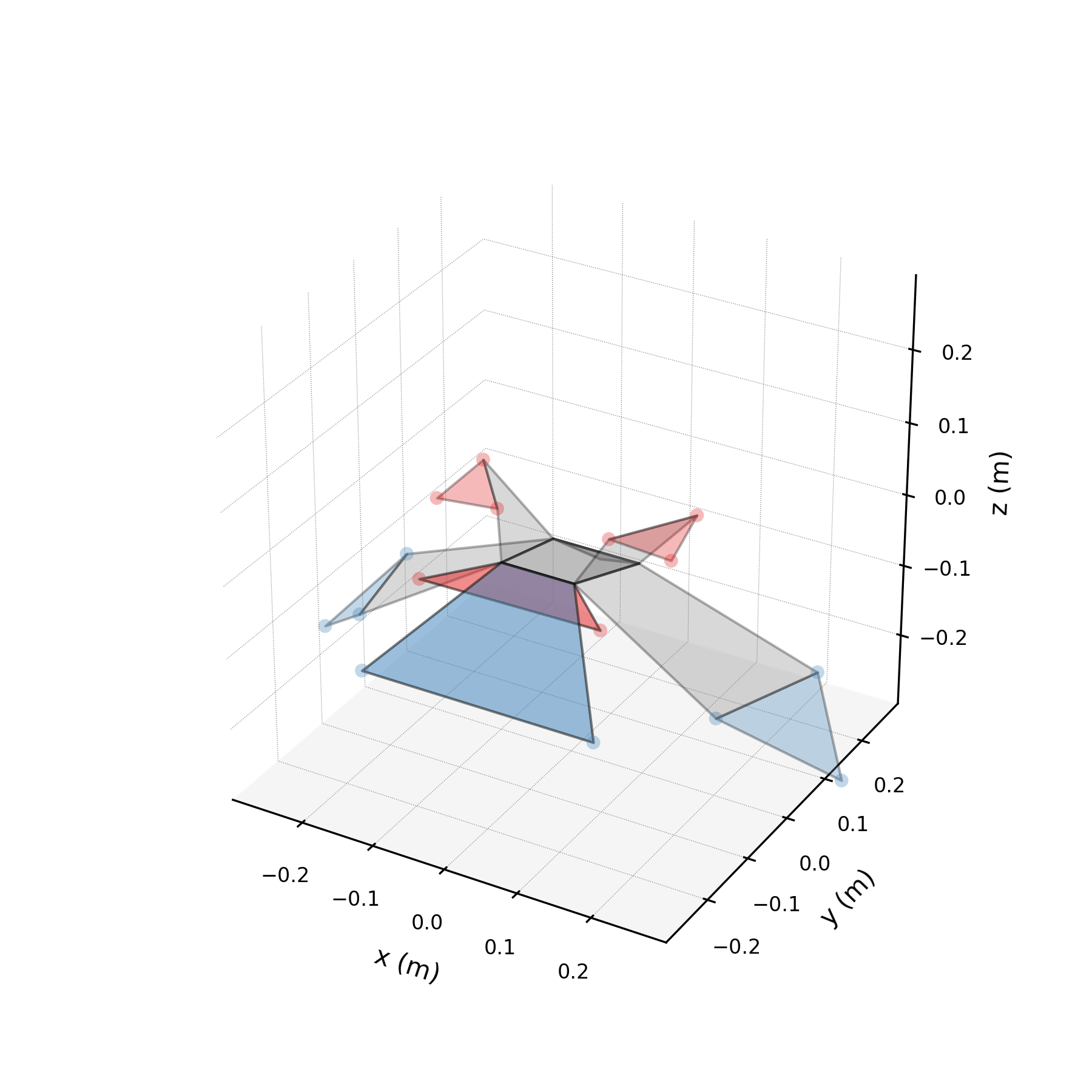
To validate the stabilised forecast over multiple wingbeat cycles, we extend the time axis to 3× the training duration and compare against the original data tiled periodically. Because the stabilised eigenvalues are purely imaginary, the forecast repeats the same spatial pattern indefinitely. Any mismatch between the forecast and tiled data reveals how well the three mode pairs capture the true periodic structure.
# Tile the original data to 3x duration for comparison with the stabilised forecast
expanded_times = expand_time_sequence(times, expansion_factor=3.0)
expanded_markers = expand_marker_sequence(times, markers, expanded_times)
# Compare wingtip z: stabilised forecast vs tiled original
fig, ax = plt.subplots(figsize=(10, 3))
ax.plot(fake_time, forecast_stable[:, 0, 2], label="Stabilised forecast", lw=1.5)
ax.plot(
expanded_times,
expanded_markers[:, 0, 2],
"--",
label="Tiled original",
lw=1,
alpha=0.7,
)
ax.axvline(times[-1], color="grey", ls=":", label="End of training data")
ax.set_xlabel("Time (s)")
ax.set_ylabel("Wingtip z (m)")
ax.legend()
ax.set_title("Extended forecast vs tiled original wingbeat")
plt.tight_layout()
fig.savefig("figures/07_extended_forecast.svg", format="svg")
plt.show()
# hawk3d_forecast = Hawk3D("../data/mean_hawk_shape.csv")
# min_len = min(forecast_stable.shape[0], expanded_markers.shape[0])
# ani = animate_compare(
# hawk3d_forecast,
# [forecast_stable[:min_len], expanded_markers[:min_len]],
# colour="blue",
# az=-60,
# el=30,
# )
# ani.save("figures/07_extended_forecast.gif", writer="Pillow", fps=40, dpi=300)
# print("Saved GIF: figures/07_extended_forecast.gif")
4 — Frequency modification¶
We can alter the wingbeat frequency while preserving the spatial mode shape. Here we reconstruct only the wingbeat pair (pair 0) at half its original frequency — stretching each stroke cycle to twice its original duration. The wing follows the same spatial path but at half speed, demonstrating that DMD cleanly separates spatial and temporal structure.
# Halve the wingbeat frequency (wingbeat pair only)
original_freq = np.abs(
np.imag(result._dmd_object.eigs[conjugate_pairs[WINGBEAT_PAIR][0]])
)
half_freq = original_freq / 2
print(f"Original wingbeat frequency: {original_freq / (2 * np.pi):.2f} Hz")
print(f"Modified wingbeat frequency: {half_freq / (2 * np.pi):.2f} Hz")
# Normal-speed wingbeat (wingbeat pair only, for comparison)
normal_wingbeat = forecast(
result,
fake_time,
pairs=[WINGBEAT_PAIR],
stable=True,
)
# Half-speed wingbeat via forecast with modified eigenvalues
slow_forecast = forecast(
result,
fake_time,
mode_indices=WINGBEAT_MODES,
stable=True,
modify_eigenvalues={
"zero_modes": WINGBEAT_MODES,
"changed_freq": half_freq,
},
)
# Compare
fig, ax = plt.subplots(figsize=(10, 3))
ax.plot(fake_time, normal_wingbeat[:, 0, 2], label="Normal wingbeat")
ax.plot(fake_time, slow_forecast[:, 0, 2], label="Half frequency")
ax.set_xlabel("Time (s)")
ax.set_ylabel("Wingtip z")
ax.legend()
ax.set_title("Frequency modification: normal vs half-speed wingbeat")
plt.tight_layout()
fig.savefig("figures/07_frequency_modification.svg", format="svg")
plt.show()
Original wingbeat frequency: 4.51 Hz Modified wingbeat frequency: 2.25 Hz
# hawk3d_up = Hawk3D("../data/mean_hawk_shape.csv")
# ani = animate_compare(
# hawk3d_up,
# [slow_forecast[:120], normal_wingbeat[:120]],
# colour="blue", az=-60, el=30,
# )
# ani.save("figures/07_half_freq_wingbeat.gif", writer='Pillow', fps=30, dpi=300)
# print("Saved GIF: figures/07_half_freq_wingbeat.gif")
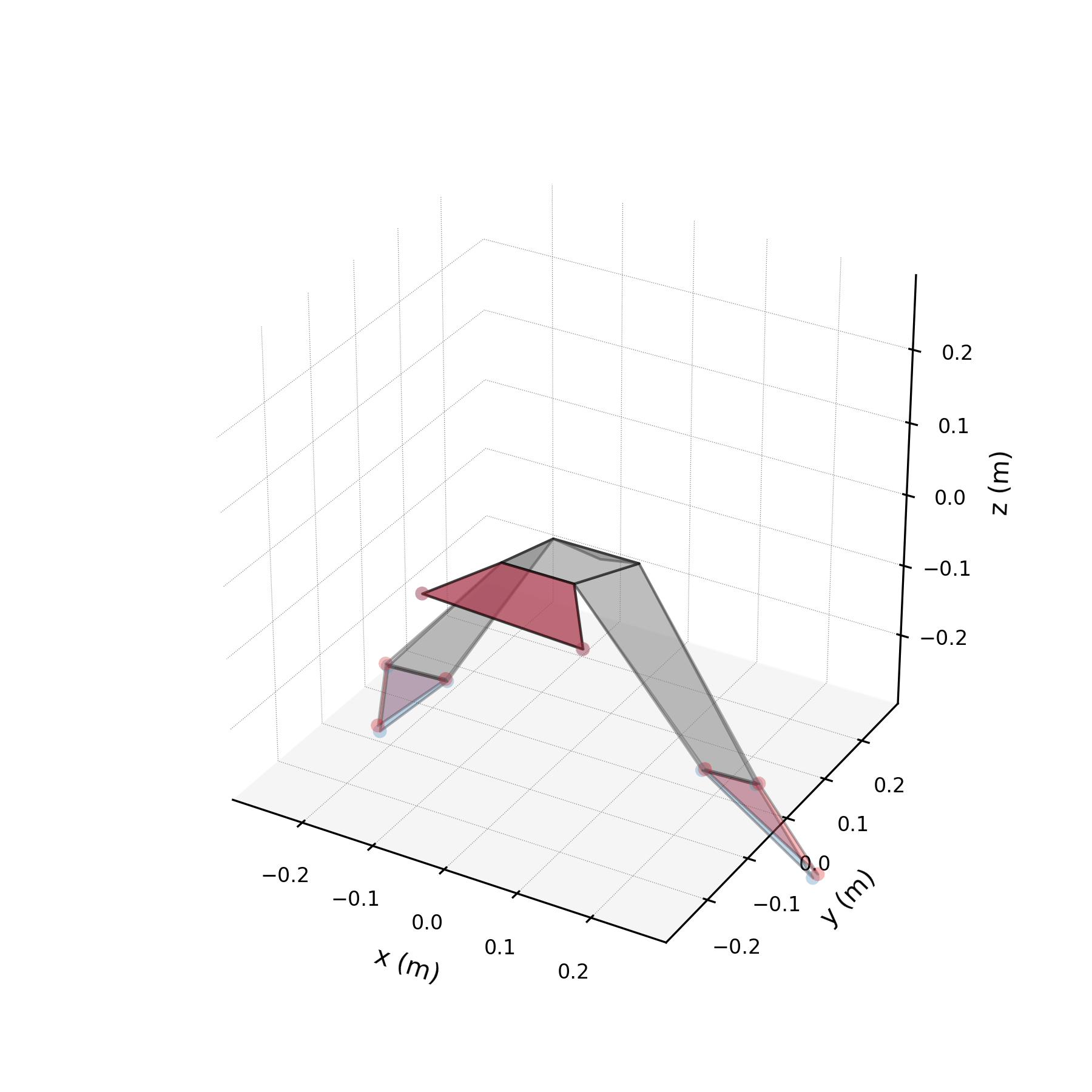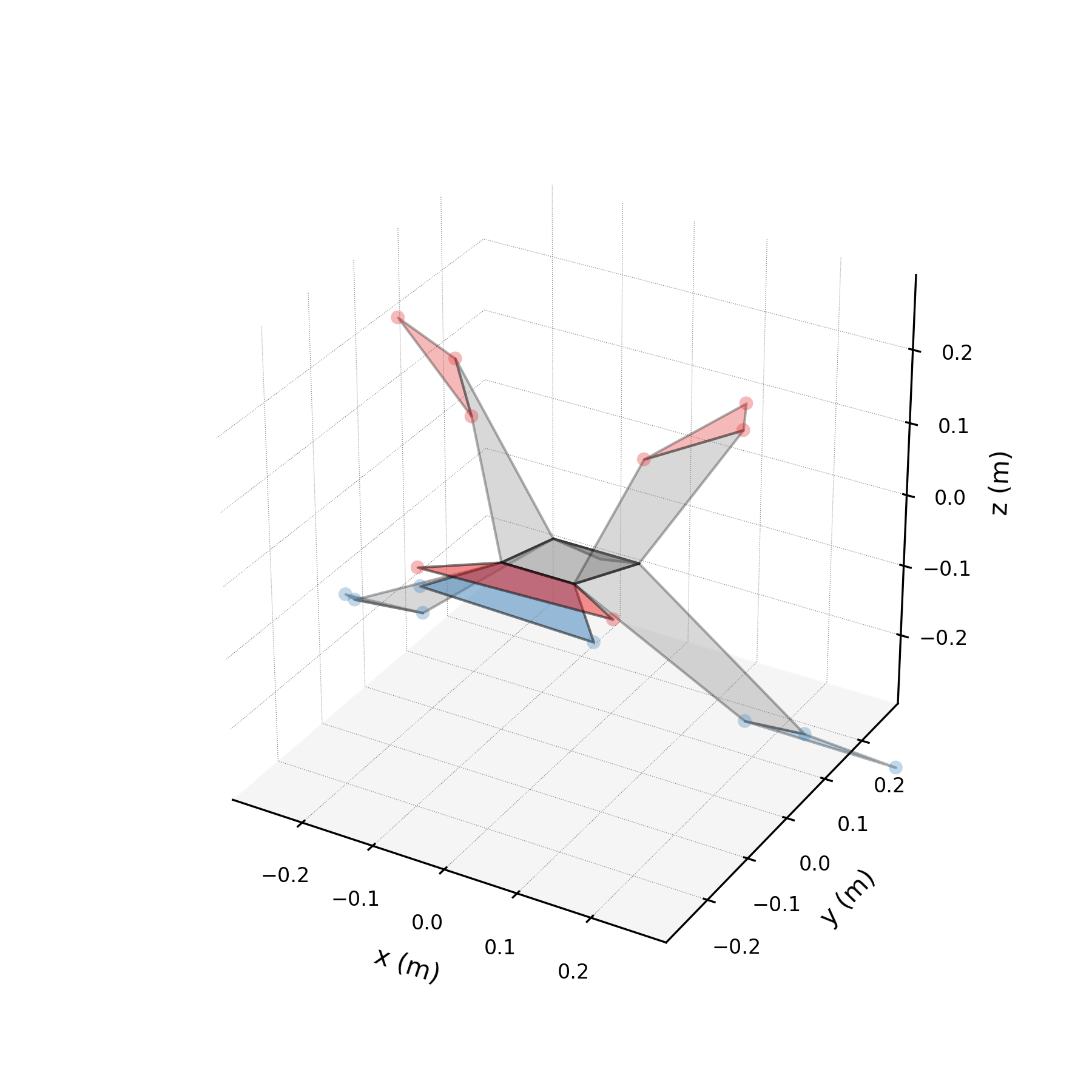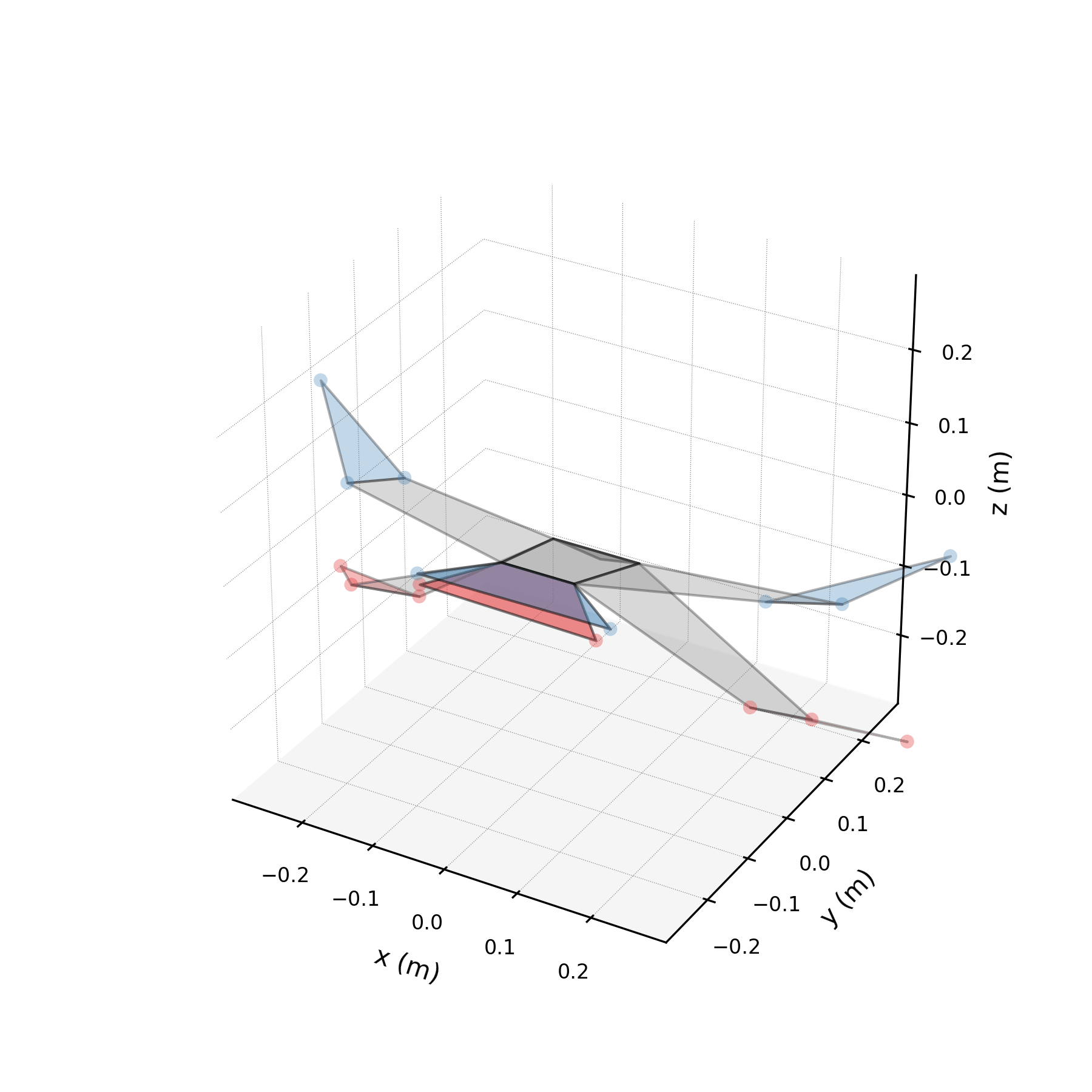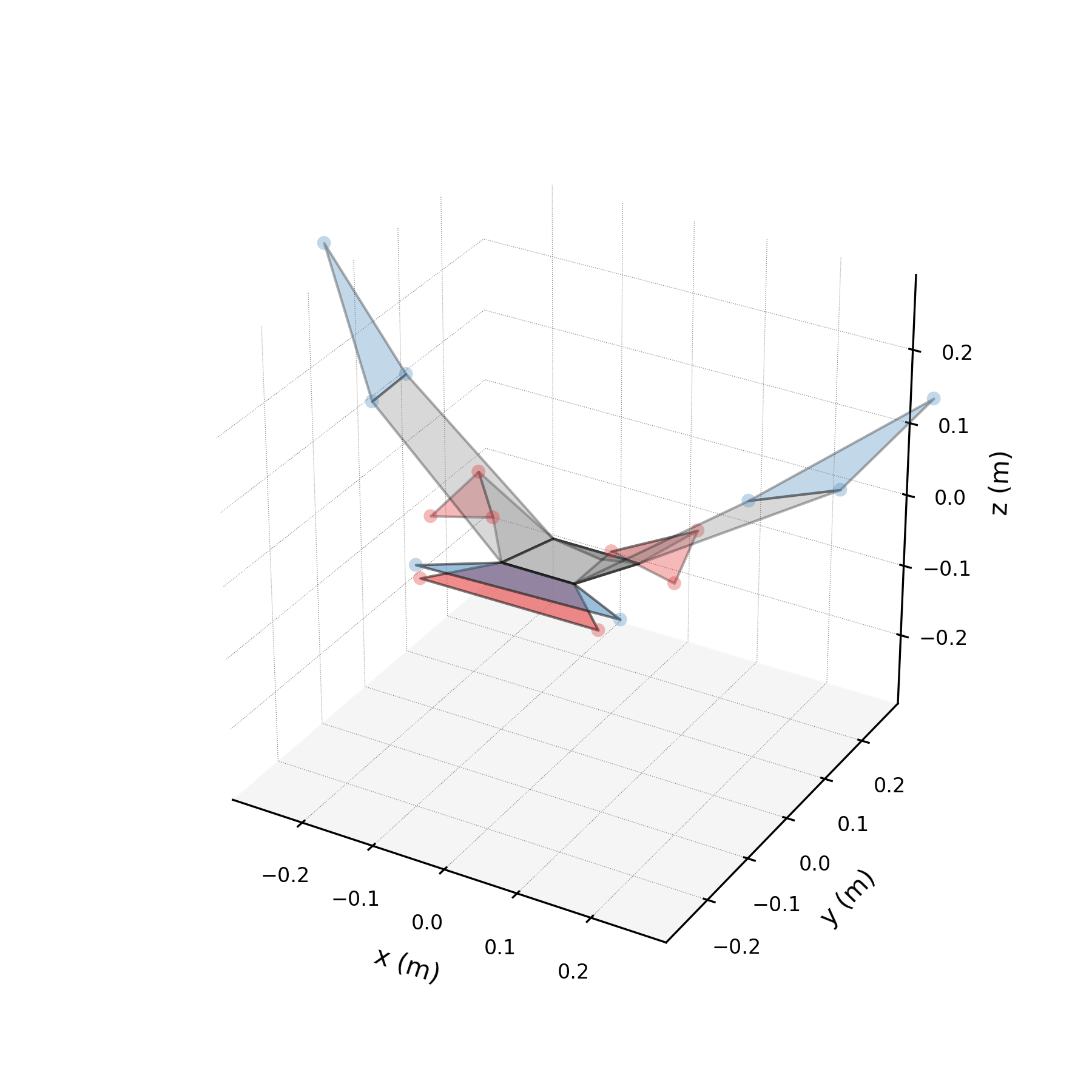
5 — Synthetic flight generation¶
Using only the wingbeat conjugate pair (stabilised), we generate a 5-second synthetic flight — far longer than any recorded sequence. Because the stabilised eigenvalues are purely imaginary, the motion repeats indefinitely without divergence.
# Generate a long synthetic flight
long_time = np.linspace(0, 5.0, 1000) # 5 seconds
synthetic_flight = forecast(
result,
long_time,
pairs=[WINGBEAT_PAIR, FOLDING_PAIR],
stable=True,
)
print(f"Synthetic flight: {synthetic_flight.shape[0]} frames, {long_time[-1]:.1f}s")
Synthetic flight: 1000 frames, 5.0s
# # Use a subset for GIF (too many frames otherwise)
# ani = animate(
# hawk3d, synthetic_flight[:200],
# colour="steelblue", az=-60, el=30,
# )
# ani.save("figures/07_synthetic_flight.gif", writer='Pillow', fps=40, dpi=300)
# print("Saved GIF: figures/07_synthetic_flight.gif")

# Remove a mode: what does flight look like without the folding mode?
no_fold = forecast(
result,
times[1:],
mode_indices=WINGBEAT_MODES + BASE_MODES, # skip folding
stable=False,
)
# Compare with full reconstruction
fig, ax = plt.subplots(figsize=(8, 3))
ax.plot(times[1:], result.reconstruction[:, 0, 2], label="Full (6 modes)")
ax.plot(times[1:], no_fold[:, 0, 2], "--", label="Without folding mode")
ax.set_xlabel("Time (s)")
ax.set_ylabel("Wingtip z")
ax.legend()
ax.set_title("Effect of removing the folding mode")
plt.tight_layout()
fig.savefig("figures/07_no_folding.svg", format="svg")
plt.show()
6 — Temporal upsampling¶
Because DMD decomposes the signal into continuous-time modes, we can evaluate the model at arbitrary time points — including a denser grid than the original data. This provides smooth temporal upsampling without requiring interpolation heuristics.
# Upsampling = evaluate the same DMD model on a denser time grid.
upsample_factor = 3
dense_time = np.linspace(times[0], times[-1], len(times) * upsample_factor)
upsampled = reconstruct(result, times=dense_time)
# Compare against original reconstruction
fig, ax = plt.subplots(figsize=(10, 3))
ax.plot(
dense_time,
upsampled[:, 0, 0],
".-",
lw=0.5,
alpha=0.5,
label=f"Upsampled ({len(dense_time)} frames)",
)
ax.plot(times, markers[:, 0, 0], "o", ms=3, label=f"Original ({len(times) - 1} frames)")
ax.set_xlabel("Time (s)")
ax.set_ylabel("Wingtip z (m)")
ax.legend()
ax.set_title(f"Temporal upsampling: {upsample_factor}\u00d7 resolution")
plt.tight_layout()
fig.savefig("figures/07_upsampled.svg", format="svg")
plt.show()
# ani = animate_compare(
# hawk3d_up,
# [upsampled[:100], np.repeat(markers, 3, axis=0)[:100]],
# colour="blue", az=-60, el=30,
# )
# ani.save("figures/07_upsampled.gif", writer='Pillow', fps=30, dpi=300)
# print("Saved GIF: figures/07_upsampled.gif")
# hawk3d_up = Hawk3D("../data/mean_hawk_shape.csv")
# animate_plotly_compare(
# hawk3d_up,
# keypoints_frames_list=[upsampled[:100], np.repeat(markers, 3, axis=0)[:100]],
# colours=["black", "blue"],
# )
Summary¶
This notebook demonstrated four generative capabilities of the DMD model: stabilised forecasting that removes exponential growth and decay, frequency modification that preserves spatial structure while changing temporal dynamics, synthetic flight generation from arbitrary mode subsets, and temporal upsampling that evaluates continuous-time modes on a denser grid. Together with the preceding notebooks, these results show that just three conjugate mode pairs — wingbeat, doubled-frequency, and base — provide both an accurate decomposition and a flexible generative model of hawk flapping flight.